📚 Academic MCP
🔬 academic-mcp is a Python-based MCP server that enables users to search, download, and read academic papers from various platforms. It provides three main tools:
- 🔎
paper_search: Search papers across multiple academic databases - 📥
paper_download: Download paper PDFs, return paths of downloaded files - 📖
paper_read: Extract and read text content from papers


📑 Table of Contents
✨ Features
- 🌐 Multi-Source Support: Search and download papers from arXiv, PubMed, bioRxiv, medRxiv, Google Scholar, IACR ePrint Archive, Semantic Scholar, and CrossRef.
- 🎯 Unified Interface: All platforms accessible through consistent
paper_search,paper_download, andpaper_readtools. - 📊 Standardized Output: Papers are returned in a consistent dictionary format via the
Paperclass. - ⚡ Asynchronous Operations: Efficiently handles concurrent searches and downloads using
httpxand async/await. - 🔌 MCP Integration: Compatible with MCP clients for LLM context enhancement.
- 🧩 Extensible Design: Easily add new academic platforms by extending the
sourcesmodule.
🎬 Screenshot
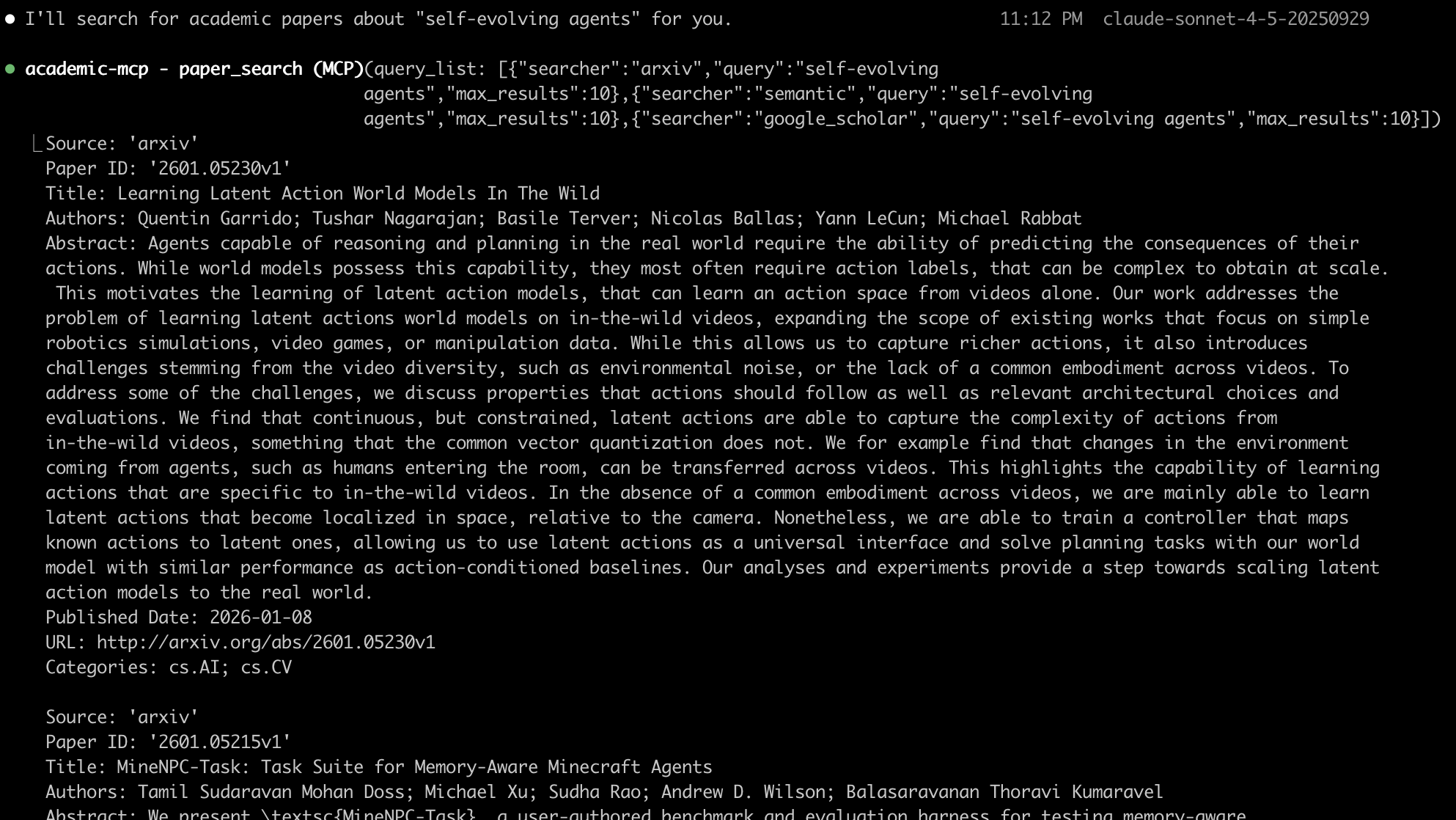
📝 TODO
Planned Academic Platforms
- arXiv
- PubMed
- bioRxiv
- medRxiv
- Google Scholar
- IACR ePrint Archive
- Semantic Scholar
- CrossRef
- PubMed Central (PMC)
- Science Direct
- Springer Link
- IEEE Xplore
- ACM Digital Library
- Web of Science
- Scopus
- JSTOR
- ResearchGate
- CORE
- Microsoft Academic
📦 Installation
academic-mcp can be installed using uv or pip. Below are two approaches: a quick start for immediate use and a detailed setup for development.
⚡ Quick Start
For users who want to quickly run the server:
-
Install Package:
pip install academic-mcp -
Configure Claude Desktop: Add this configuration to
~/Library/Application Support/Claude/claude_desktop_config.json(Mac) or%APPDATA%\Claude\claude_desktop_config.json(Windows):{ "mcpServers": { "academic-mcp": { "command": "python", "args": [ "-m", "academic_mcp" ], "env": { "SEMANTIC_SCHOLAR_API_KEY": "", "ACADEMIC_MCP_DOWNLOAD_PATH": "./downloads" } } } }Note: The
SEMANTIC_SCHOLAR_API_KEYis optional and only required for enhanced Semantic Scholar features.
🛠️ For Development
For developers who want to modify the code or contribute:
-
Setup Environment:
# Install uv if not installed curl -LsSf https://astral.sh/uv/install.sh | sh # Clone repository git clone https://github.com/LinXueyuanStdio/academic-mcp.git cd academic-mcp # Create and activate virtual environment uv venv source .venv/bin/activate # On Windows: .venv\Scripts\activate -
Install Dependencies:
# Install dependencies (recommended) uv pip install -e . # Add development dependencies (optional) uv pip install pytest flake8
🚀 Usage
Once configured, academic-mcp provides three main tools accessible through Claude Desktop or any MCP-compatible client:
1. Search Papers (paper_search)
Search for academic papers across multiple sources:
# Search arXiv for machine learning papers
paper_search([
{"searcher": "arxiv", "query": "machine learning", "max_results": 5}
])
# Search multiple platforms simultaneously
paper_search([
{"searcher": "arxiv", "query": "deep learning", "max_results": 5},
{"searcher": "pubmed", "query": "cancer immunotherapy", "max_results": 3},
{"searcher": "semantic", "query": "climate change", "max_results": 4, "year": "2020-2023"}
])
# Search all platforms (omit "searcher" parameter)
paper_search([
{"query": "quantum computing", "max_results": 10}
])
2. Download Papers (paper_download)
Download paper PDFs using their identifiers:
paper_download([
{"searcher": "arxiv", "paper_id": "2106.12345"},
{"searcher": "pubmed", "paper_id": "32790614"},
{"searcher": "biorxiv", "paper_id": "10.1101/2020.01.01.123456"},
{"searcher": "semantic", "paper_id": "DOI:10.18653/v1/N18-3011"}
])
3. Read Papers (paper_read)
Extract and read text content from papers:
# Read an arXiv paper
paper_read(searcher="arxiv", paper_id="2106.12345")
# Read a PubMed paper
paper_read(searcher="pubmed", paper_id="32790614")
# Read a Semantic Scholar paper
paper_read(searcher="semantic", paper_id="DOI:10.18653/v1/N18-3011")
Environment Variables
SEMANTIC_SCHOLAR_API_KEY: Optional API key for enhanced Semantic Scholar featuresACADEMIC_MCP_DOWNLOAD_PATH: Directory for downloaded PDFs (default:./downloads)
🤝 Contributing
We welcome contributions! Here's how to get started:
-
Fork the Repository: Click "Fork" on GitHub.
-
Clone and Set Up:
git clone https://github.com/yourusername/academic-mcp.git cd academic-mcp uv pip install -e . # Install in development mode -
Make Changes:
- Add new platforms in
academic_mcp/sources/. - Update tests in
tests/.
- Add new platforms in
-
Submit a Pull Request: Push changes and create a PR on GitHub.
📄 License
This project is licensed under the MIT License. See the LICENSE file for details.
Happy researching with academic-mcp! If you encounter issues, open a GitHub issue.
