Pedigree MCP Server
An MCP (Model Context Protocol) server for generating family pedigree tree diagrams as PNG images using standard genetic notation per Bennett 2008/2022 NSGC guidelines.
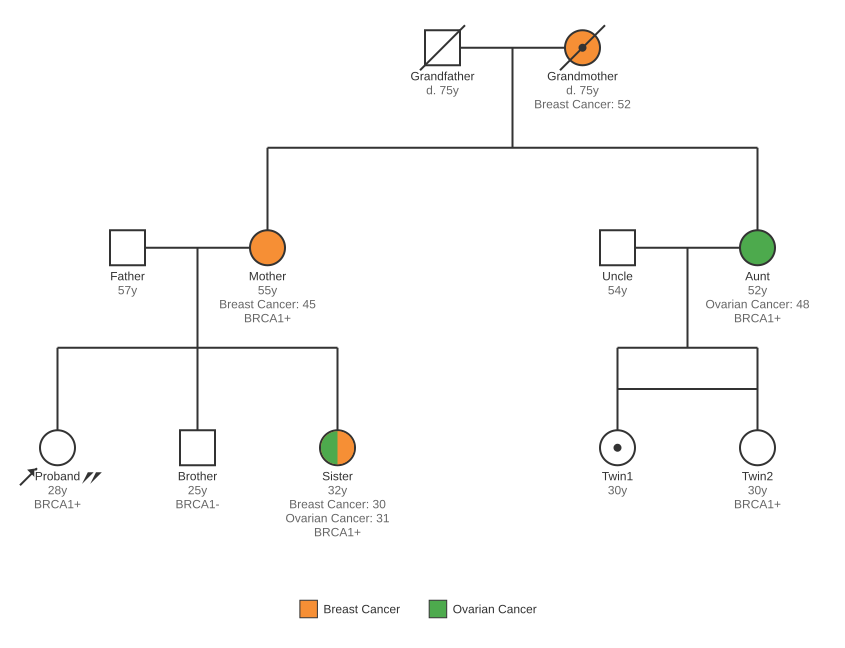
BRCA1 hereditary breast/ovarian cancer family history demonstrating: 3 generations, affected individuals with combined conditions, MZ identical twins, carrier status, deceased indicators, proband + consultand markers, and gene test results (BRCA1+/BRCA1-)
Installation
Add to your MCP client configuration (e.g., Claude Desktop):
{
"mcpServers": {
"pedigree": {
"command": "npx",
"args": ["pedigree-mcp"],
"env": {}
}
}
}
Build from Source
If you prefer to build from source:
git clone https://github.com/zzgael/pedigree-mcp.git
cd pedigree-mcp
npm install
npm run build
Then use in your MCP client configuration:
{
"mcpServers": {
"pedigree": {
"command": "node",
"args": ["/absolute/path/to/pedigree-mcp/dist/index.js"],
"env": {}
}
}
}
Features
Bennett 2008 Standard Compliance
This implementation follows the NSGC Standardized Human Pedigree Nomenclature:
| Symbol | Description | Property |
|---|---|---|
| Square | Male | sex: "M" |
| Circle | Female | sex: "F" |
| Diamond | Unknown sex | sex: "U" |
| Filled shape | Affected individual | conditions: [...] |
| Diagonal line | Deceased | status: 1 |
| Arrow (lower-left) | Proband | proband: true |
| Double arrow (lower-left) | Consultand (person seeking counseling) | consultand: true |
| Brackets [ ] | Adopted | noparents: true |
| Double line | Consanguinity | Auto-detected from shared ancestors |
| Text on double line | Consanguinity degree | consanguinity_degree: "1st cousins" |
| Horizontal bar | MZ (identical) twins | mztwin: "group_id" |
| Diagonal lines | DZ (fraternal) twins | dztwin: "group_id" |
| Dot in center | Carrier status | carrier: true |
| Outlined dot | Obligate carrier (inferred) | obligate_carrier: true |
| "P" inside symbol | Pregnancy | pregnant: true |
| "P" + weeks label | Pregnancy duration | pregnant: true, terminated_age: 12 |
| Small triangle | Early pregnancy loss (<20 weeks) | terminated: true, terminated_age: 8 |
| Large triangle | Stillbirth (≥20 weeks) | terminated: true, terminated_age: 24 |
| "EP" below symbol | Ectopic pregnancy | ectopic: true |
| Crossed lines (X) | Infertility | infertility: true |
| Hash marks on line | Divorced/separated | divorced: true |
| Line through offspring | No children by choice | no_children_by_choice: true |
| "A" in upper right | Ashkenazi ancestry | ashkenazi: 1 |
| "*" in upper left | Genetic anticipation | anticipation: true |
| "d. XXy" label | Age at death (auto-calculated) | yob: 1950, yod: 2020, status: 1 |
| Arrow + "OUT" label | Adopted OUT (placed for adoption) | adoption_type: "out" |
| Dashed brackets | Foster placement (temporary) | adoption_type: "foster" |
| Roman numerals I, II, III | Birth order in sibling group | birth_order: 1 (displays as "I") |
| "E" marker (blue) | ART - Egg donor conception | art_type: "egg_donor" |
| "S" marker (blue) | ART - Sperm donor conception | art_type: "sperm_donor" |
| "Em" marker (blue) | ART - Embryo donor conception | art_type: "embryo_donor" |
| "GC" marker (blue) | ART - Gestational carrier (surrogate) | art_type: "surrogate" |
| "SAB" label | Pregnancy outcome - Spontaneous abortion | pregnancy_outcome: "miscarriage" |
| "TOP" label | Pregnancy outcome - Termination of pregnancy | pregnancy_outcome: "induced_termination" |
| "SB" label | Pregnancy outcome - Stillbirth | pregnancy_outcome: "stillbirth" |
| "Het" label (green) | Gene copy number - Heterozygous | gene_copy_number: "heterozygous" |
| "Hom" label (green) | Gene copy number - Homozygous | gene_copy_number: "homozygous" |
| "CH" label (green) | Gene copy number - Compound heterozygous | gene_copy_number: "compound_heterozygous" |
| Dashed partnership line | Unmarried/common-law partnership | relationship_type: "unmarried" |
Conditions (Bennett Standard - FREE TEXT)
Per Bennett 2008 standard, conditions are documented using free text. Simply provide a conditions array with any disease/condition name:
{
"conditions": [
{ "name": "Breast cancer", "age": 42 },
{ "name": "Ovarian cancer", "age": 55 }
]
}
Examples:
{ "name": "Huntington's disease", "age": 45 }{ "name": "Type 2 diabetes" }(no age = affected status only){ "name": "Cystic fibrosis" }{ "name": "Hereditary hemochromatosis", "age": 38 }
Colors are auto-assigned from a palette based on unique condition names. Multiple conditions show as quadrants (male) or pie slices (female).
Genetic Testing Results
Supports any gene - use pattern {gene}_gene_test:
{
"brca1_gene_test": { "type": "T", "result": "P" },
"htt_gene_test": { "type": "T", "result": "P" },
"apoe_gene_test": { "type": "S", "result": "N" }
}
Gene test result codes:
- type:
T(tested),S(screening),-(unknown) - result:
P(positive),N(negative),-(unknown/VUS)
Labels appear as: BRCA1+ (positive), HTT- (negative)
Tools
get_pedigree_documentation
Returns comprehensive documentation about the pedigree data format. Always call this first before generating a pedigree.
generate_pedigree
Generates a family pedigree tree in PNG or SVG format.
Parameters:
| Parameter | Type | Default | Description |
|---|---|---|---|
dataset | Individual[] | required | Array of family members |
width | number | 800 | Image width in pixels |
height | number | 600 | Image height in pixels |
symbol_size | number | 35 | Node diameter in pixels |
background | string | #ffffff | Background color |
labels | string[] | ['age'] | Attributes to display |
format | 'png' | 'svg' | 'png' | Output format: png (base64 image) or svg (XML text) |
Data Format
Individual Object
interface Individual {
// Required
name: string; // Unique ID (max 7 chars)
sex: "M" | "F" | "U"; // Male, Female, Unknown
// Identity
display_name?: string; // Human-readable name for display (max 13 chars)
top_level?: boolean; // Founding individual (no parents)
proband?: boolean; // Index case
// Relationships
mother?: string; // Mother's name (must exist in dataset)
father?: string; // Father's name (must exist in dataset)
// Demographics
age?: number; // Current age
yob?: number; // Year of birth
status?: number; // 0 = alive, 1 = deceased
// Twins (Bennett standard)
mztwin?: string; // MZ twin group ID (identical)
dztwin?: string; // DZ twin group ID (fraternal)
// Special indicators (Bennett standard)
carrier?: boolean; // Carrier status (dot in center)
pregnant?: boolean; // Current pregnancy (P inside symbol)
terminated?: boolean; // Stillbirth/SAB (small triangle)
divorced?: boolean; // Divorced from partner (hash marks)
noparents?: boolean; // Adopted (brackets around symbol)
// Conditions (Bennett standard - FREE TEXT)
conditions?: Array<{
name: string; // Any condition: "Breast cancer", "Huntington's disease", etc.
age?: number; // Age at diagnosis/onset
}>;
// Genetic tests (pattern: {gene}_gene_test)
brca1_gene_test?: { type: "-"|"S"|"T", result: "-"|"P"|"N" };
brca2_gene_test?: { type: "-"|"S"|"T", result: "-"|"P"|"N" };
// ... any gene test
}
Examples
📸 View All 21 Scenario Examples →
See the full gallery of standardized pedigree scenarios demonstrating Bennett 2008/2022 compliance, including gender diversity, twins, consanguinity, ART indicators, and complex multi-generation families.
Basic Three-Generation Pedigree
[
{"name": "MGF", "sex": "M", "top_level": true},
{"name": "MGM", "sex": "F", "top_level": true, "conditions": [{"name": "Breast cancer", "age": 55}]},
{"name": "Mother", "sex": "F", "mother": "MGM", "father": "MGF", "conditions": [{"name": "Breast cancer", "age": 42}]},
{"name": "Father", "sex": "M", "top_level": true},
{"name": "Proband", "display_name": "Sarah", "sex": "F", "mother": "Mother", "father": "Father", "proband": true, "age": 25, "brca1_gene_test": {"type": "T", "result": "P"}}
]
Neurological Condition Pedigree
[
{"name": "GF", "sex": "M", "top_level": true, "status": 1, "conditions": [{"name": "Huntington's disease", "age": 52}]},
{"name": "GM", "sex": "F", "top_level": true},
{"name": "Father", "sex": "M", "mother": "GM", "father": "GF", "conditions": [{"name": "Huntington's disease", "age": 48}]},
{"name": "Mother", "sex": "F", "top_level": true},
{"name": "Proband", "sex": "M", "mother": "Mother", "father": "Father", "proband": true, "age": 25, "carrier": true}
]
Twins Example
[
{"name": "Dad", "sex": "M", "top_level": true},
{"name": "Mom", "sex": "F", "top_level": true},
{"name": "Twin1", "sex": "M", "mother": "Mom", "father": "Dad", "mztwin": "mz1"},
{"name": "Twin2", "sex": "M", "mother": "Mom", "father": "Dad", "mztwin": "mz1"},
{"name": "DZTwin1", "sex": "M", "mother": "Mom", "father": "Dad", "dztwin": "dz1"},
{"name": "DZTwin2", "sex": "F", "mother": "Mom", "father": "Dad", "dztwin": "dz1"}
]
Complex Family with Bennett Features
[
{"name": "GF", "sex": "M", "top_level": true, "status": 1},
{"name": "GM", "sex": "F", "top_level": true, "carrier": true},
{"name": "Father", "sex": "M", "mother": "GM", "father": "GF", "divorced": true},
{"name": "Mother", "sex": "F", "top_level": true},
{"name": "Child1", "sex": "F", "mother": "Mother", "father": "Father", "proband": true, "noparents": true},
{"name": "Loss", "sex": "U", "mother": "Mother", "father": "Father", "terminated": true}
]
Development
# Install dependencies
npm install
# Run in development mode (watch)
npm run dev
# Run all tests
npm test
# Build for production
npm run build
# Type check
npx tsc --noEmit
Testing
- 159 tests total covering:
- Validation (parent references, gender constraints)
- SVG rendering (all symbol types, indicators)
- Condition markers and multi-condition pie charts
- Gene test formatting
- Twin rendering (MZ with bar, DZ without)
- Consanguinity detection
- Bennett 2008 compliance (carrier, pregnancy, termination, divorced)
- Edge cases (deep pedigrees, wide generations, half-siblings)
References
- Bennett et al. 2008 - Standardized Human Pedigree Nomenclature
- Bennett et al. 2022 - Sex and Gender Inclusivity Update
- Iowa Human Genetics - How to Draw a Pedigree
License
MIT License - see LICENSE



