SciTeX (scitex)
Python Library for Science. For AI and Human Researchers
Docs ·
Quick Start ·
API ·
pip install scitex[all]
This repository provides scitex, the orchestration layer of the SciTeX ecosystem — solving key problems in scientific research:
Problem and Solution
| # | Problem | Solution |
|---|---|---|
| 1 | Fragmented tools -- literature search, statistics, figures, and writing each require separate tools with incompatible formats | Unified toolkit -- import scitex as stx provides 50+ modules under one namespace, accessible via Python API, CLI, and MCP |
| 2 | No verification -- existing tools address whether work could be reproduced, not whether it has been verified | Cryptographic verification -- Clew builds SHA-256 hash-chain DAGs linking every manuscript claim back to source data |
| 3 | AI agents lack context -- general-purpose LLMs cannot operate across the full research lifecycle without domain-specific tools | 293 MCP tools -- AI agents run statistics, create figures, search literature, and compile manuscripts through structured tool calls |
| 4 | No custom tooling -- every lab needs domain-specific tools, but building and sharing them requires deep infrastructure knowledge | App Maker and Store -- researchers create custom apps with scitex-app SDK and share via SciTeX Cloud |
SciTeX and Research Workflow
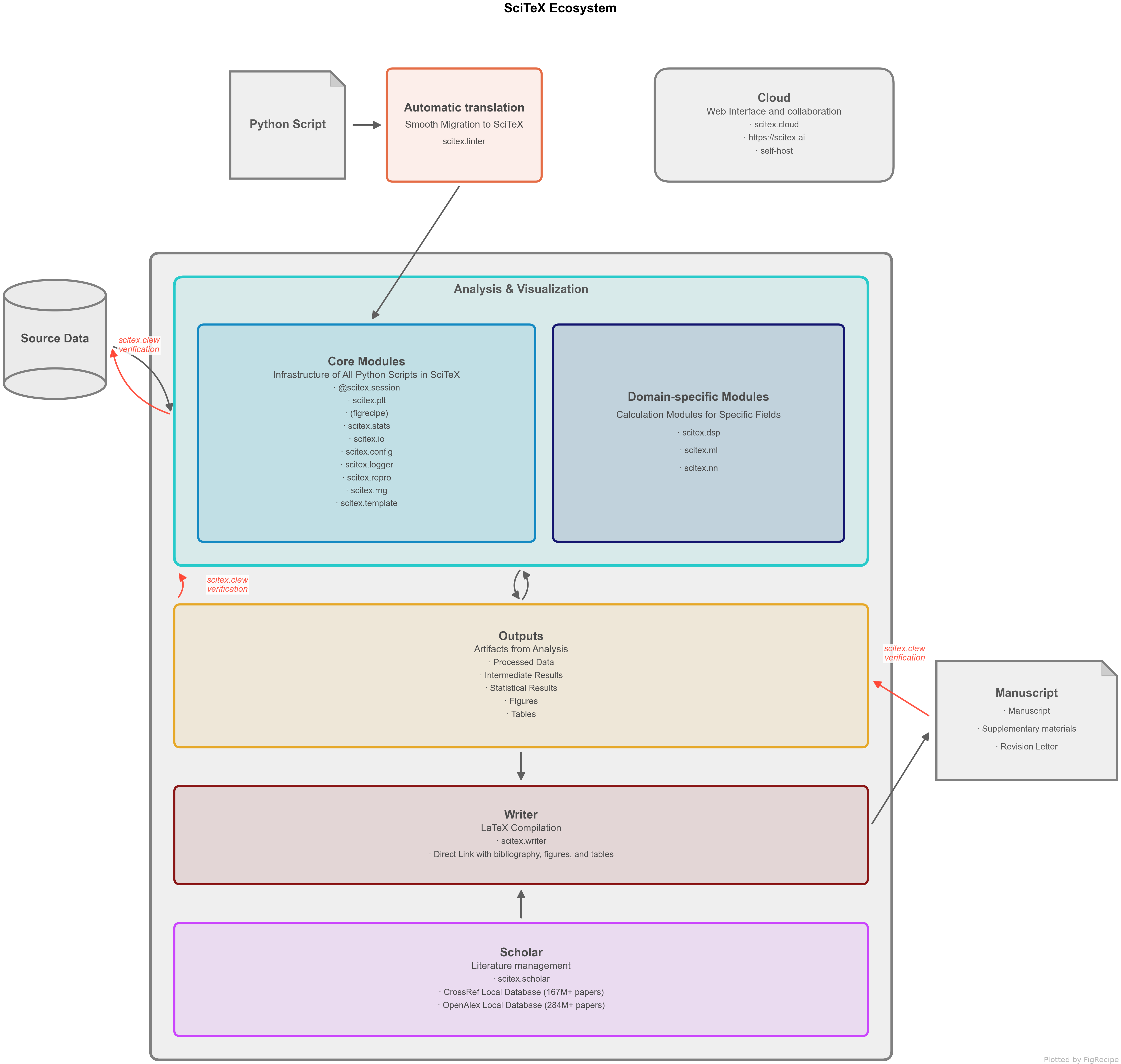
Figure 1. SciTeX research pipeline -- from literature search to manuscript compilation, with every step cryptographically linked.
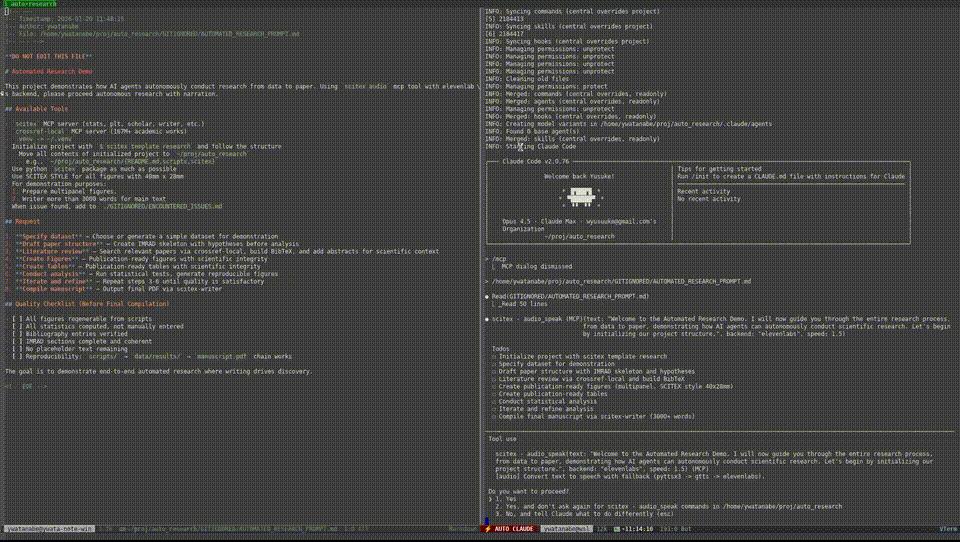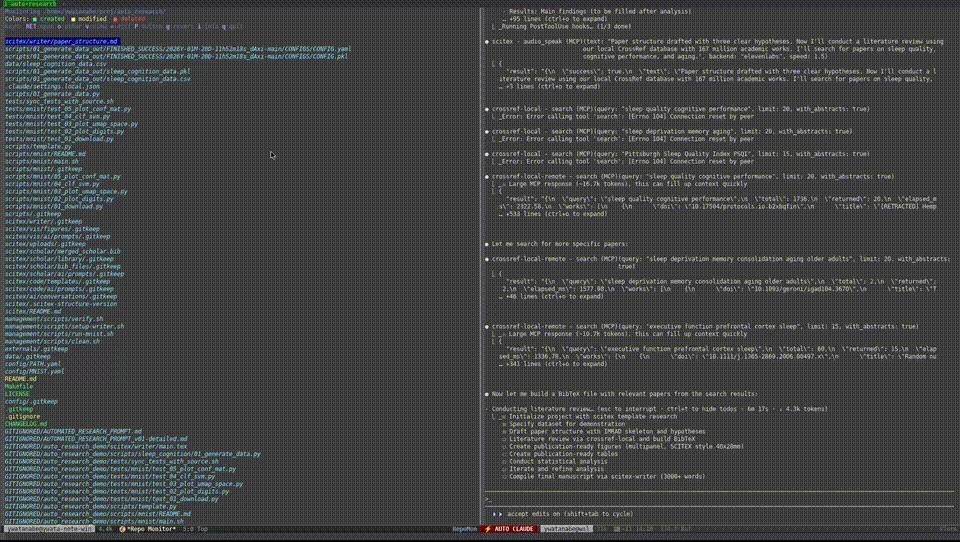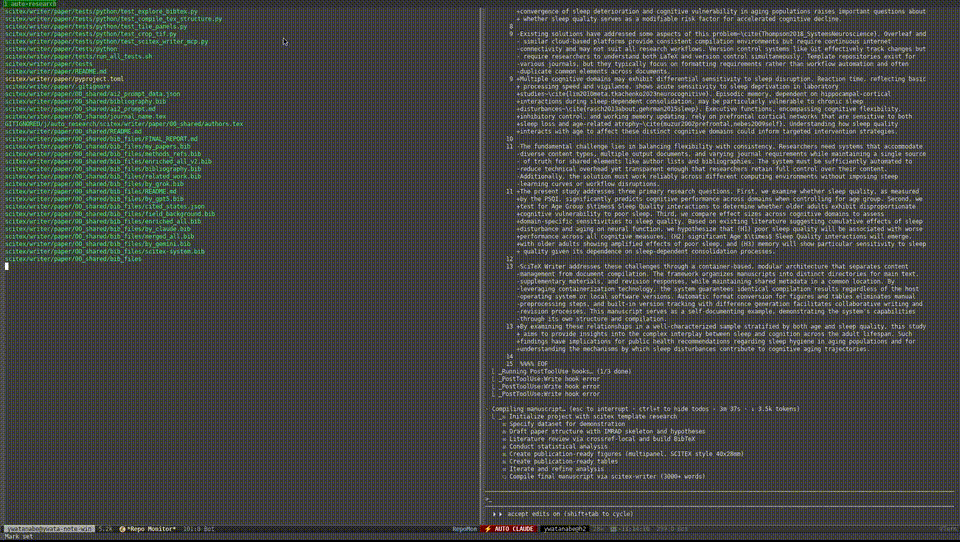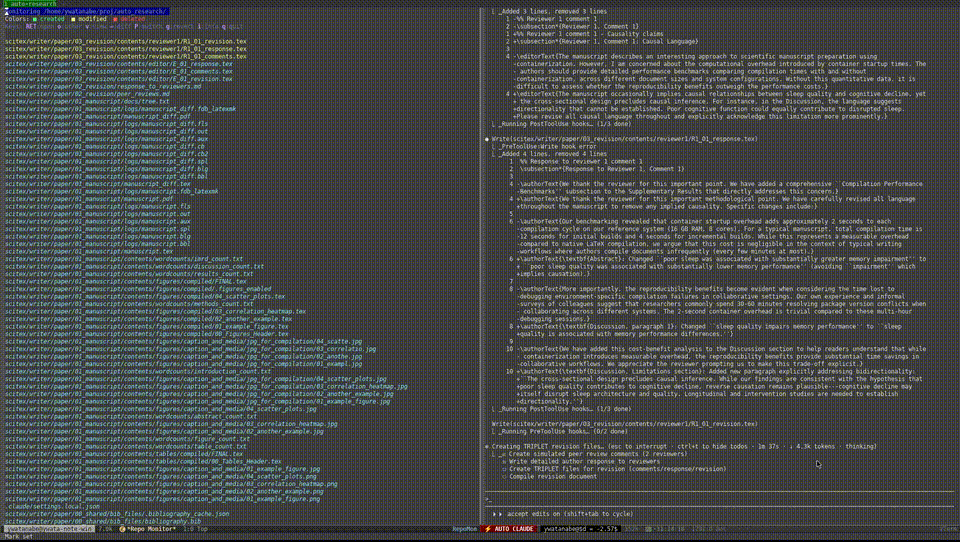
Demo — Automated Research from Data to Manuscript
40 min, minimal human intervention — an AI agent using SciTeX completed a full research cycle: literature search, statistical analysis, publication-ready figures, a 21-page manuscript, and peer review simulation. More demos are available at https://scitex.ai/demos/.
Installation
pip install scitex[all] # Recommended: everything
Per-module extras
pip install scitex # Core only (minimal)
pip install scitex[plt,stats,scholar] # Typical research setup
pip install scitex[plt] # Publication-ready figures (figrecipe)
pip install scitex[stats] # Statistical testing (23+ tests)
pip install scitex[scholar] # Literature search, PDF download, BibTeX enrichment
pip install scitex[writer] # LaTeX manuscript compilation
pip install scitex[audio] # Text-to-speech
pip install scitex[ai] # LLM APIs (OpenAI, Anthropic, Google) + ML tools
pip install scitex[dataset] # Scientific datasets (DANDI, OpenNeuro, PhysioNet)
pip install scitex[browser] # Web automation (Playwright)
pip install scitex[capture] # Screenshot capture and monitoring
pip install scitex[cloud] # Cloud platform integration
Requires Python 3.10+. We recommend uv for fast installs.
Module Overview
| Category | Modules | Description |
|---|---|---|
| Core | session, io, config, clew | Experiment tracking, file I/O, config, cryptographic verification |
| Analysis | stats, plt, dsp, linalg | Statistics, plotting, signal processing, linear algebra |
| Research | scholar, writer, diagram, canvas | Literature, manuscripts, diagrams, figure composition |
| ML/AI | ai, nn, torch, cv, benchmark | LLM APIs, neural networks, PyTorch, computer vision |
| Data | pd, db, dataset, schema | Pandas utilities, databases, scientific datasets |
| Infra | app, cloud, tunnel, container | App SDK, cloud, SSH tunnels, containers |
| Automation | browser, capture, audio, notification | Web automation, screenshots, TTS, notifications |
| Dev | dev, template, linter, introspect | Ecosystem tools, scaffolding, code analysis |
Quick Start
@scitex.session -- Reproducible Experiment Tracking
One decorator gives you: auto-CLI, YAML config injection, random seed fixation, structured output, and logging.
import scitex as stx
import numpy as np
@stx.session
def main(
data_path: str = "./data.csv", # --data-path data.csv
n_samples: int = 100, # --n-samples 200
CONFIG=stx.session.INJECTED, # Aggregated ./config/*.yaml
plt=stx.session.INJECTED, # Pre-configured matplotlib
logger=stx.session.INJECTED, # Session logger
):
"""Analyze data. Docstring becomes --help text."""
# Load
data = stx.io.load(data_path)
# Demo data
x = np.linspace(0, 2 * np.pi, n_samples)
y = np.sin(x) + np.random.randn(n_samples) * 0.1
# FigRecipe Plot
fig, ax = stx.plt.subplots()
ax.plot(x, y)
ax.set_xyt("Time", "Amplitude", "Noisy Sine Wave")
# Save sine.png + sine.csv with logging message
stx.io.save(fig, "sine.png")
return 0
if __name__ == "__main__":
main()
$ python script.py --data-path experiment.csv --n-samples 200
$ python script.py --help
# usage: script.py [-h] [--data-path DATA_PATH] [--n-samples N_SAMPLES]
# Analyze data. Docstring becomes --help text.
script_out/FINISHED_SUCCESS/2026-03-18_14-30-00_Z5MR/
├── sine.png, sine.csv # Figure + auto-exported plot data
├── CONFIGS/CONFIG.yaml # Frozen parameters
└── logs/{stdout,stderr}.log # Execution logs
scitex.io -- Unified File I/O (50+ Formats)
import scitex as stx
# Save and load -- format detected from extension
stx.io.save(df, "results.csv")
df = stx.io.load("results.csv")
stx.io.save(arr, "data.npy")
arr = stx.io.load("data.npy")
stx.io.save(fig, "figure.png") # Also exports figure data as CSV
stx.io.save(config, "config.yaml")
stx.io.save(model, "model.pkl")
# Aggregate ./config/*.yaml into a single DotDict
CONFIG = stx.io.load_configs(config_dir="./config")
print(CONFIG.MODEL.hidden_size) # Dot-notation access
# Register custom formats
@stx.io.register_saver(".custom")
def save_custom(obj, path, **kw):
with open(path, "w") as f:
f.write(str(obj))
@stx.io.register_loader(".custom")
def load_custom(path, **kw):
with open(path) as f:
return f.read()
Supports: CSV, JSON, YAML, TOML, HDF5, NPY, NPZ, PKL, PNG, JPG, SVG, PDF, Excel, Parquet, Zarr, INI, TXT, MAT, WAV, MP3, BibTeX, and more.
Built-in features: Auto directory creation, path resolution to <script_name>_out/, symlinks (symlink_from_cwd=True), save logging with file size, and Clew hash tracking.
scitex.plt -- Reproducible, Restylable Figures
Powered by figrecipe. Figures are reproducible nodes in the Clew verification DAG -- scientific data and visual style are decomposed, so figures can be restyled (fonts, colors, layout) without altering the underlying data hash. Every figure auto-exports its data as CSV + a YAML recipe for exact reproduction.
import scitex as stx
fig, axes = stx.plt.subplots(1, 3)
axes[0].stx_line(x, y)
axes[0].set_xyt("Time", "Value", "Line")
axes[1].stx_violin([g1, g2, g3])
axes[1].set_xyt("Group", "Score", "Violin")
axes[2].stx_heatmap(corr_matrix)
axes[2].set_xyt("X", "Y", "Heatmap")
stx.io.save(fig, "analysis.png") # Saves analysis.png + analysis.csv + analysis.yaml
# Restyle without changing data (hash stays valid for Clew verification)
stx.plt.reproduce("analysis.yaml", style="nature")
scitex.stats -- Publication-Ready Statistics (23+ Tests)
import scitex as stx
result = stx.stats.run_test("ttest_ind", group1, group2, return_as="dataframe")
# Returns: p-value, effect size (Cohen's d), CI, normality check, power
recommendations = stx.stats.recommend_tests(data)
stx.stats.format_results(result, style="apa") # "t(58) = 2.34, p = .021, d = 0.60"
scitex.scholar -- Literature Management
Search, download, enrich papers. Backed by local CrossRef (167M+) and OpenAlex (250M+) databases.
import scitex as stx
papers = stx.scholar.search("neural oscillations working memory", n=20)
stx.scholar.fetch("10.1038/s41586-024-07804-3")
stx.scholar.enrich_bibtex("references.bib", output="enriched.bib")
scitex scholar search "neural oscillations" --n 20
scitex scholar bibtex references.bib --output enriched.bib
scitex.writer -- LaTeX Manuscript Compilation
import scitex as stx
stx.writer.compile_manuscript("paper/")
stx.writer.add_figure("paper/", "results.png", caption="Main results")
stx.writer.add_table("paper/", "stats.csv", caption="Statistical summary")
scitex.notification -- Multi-Backend Notifications
Get notified when experiments finish -- via desktop, phone call, SMS, or email -- with automatic fallback.
import scitex as stx
stx.notification.alert("Experiment complete: accuracy = 94.2%")
stx.notification.call("Training diverged -- loss is NaN")
stx.notification.sms("GPU job finished on node-42")
@stx.session(notify=True) # Notifies on completion or failure
def main(CONFIG=stx.session.INJECTED): ...
scitex.clew -- Cryptographic Verification for AI-Driven Science
As AI agents produce research at scale, the question shifts from "could this be reproduced?" to "has this been verified?". Clew builds a SHA-256 hash-chain DAG linking every manuscript claim back to source data.
import scitex as stx
# Every stx.io.load/save automatically records file hashes -- zero config
stx.clew.status() # {'verified': 12, 'mismatched': 0, 'missing': 0}
stx.clew.chain("results/figure1.png") # Trace one file back to source data
stx.clew.dag(claims=True) # Verify all manuscript claims
# Register traceable assertions
stx.clew.add_claim(
file_path="paper/main.tex", claim_type="statistic", line_number=142,
claim_value="t(58) = 2.34, p = .021",
source_session="2026-03-18_14-30-00_Z5MR", source_file="results/stats.csv",
)
stx.clew.mermaid(claims=True) # Visualize provenance DAG
| Mode | Function | Answers |
|---|---|---|
| Project | clew.dag() | Is the whole project intact? |
| File | clew.chain("output.csv") | Can I trust this specific file? |
| Claim | clew.verify_claim("Fig 1") | Is this manuscript assertion valid? |
L1 hash comparison (ms) / L2 sandbox re-execution (min) / L3 registered timestamp proof (optional).
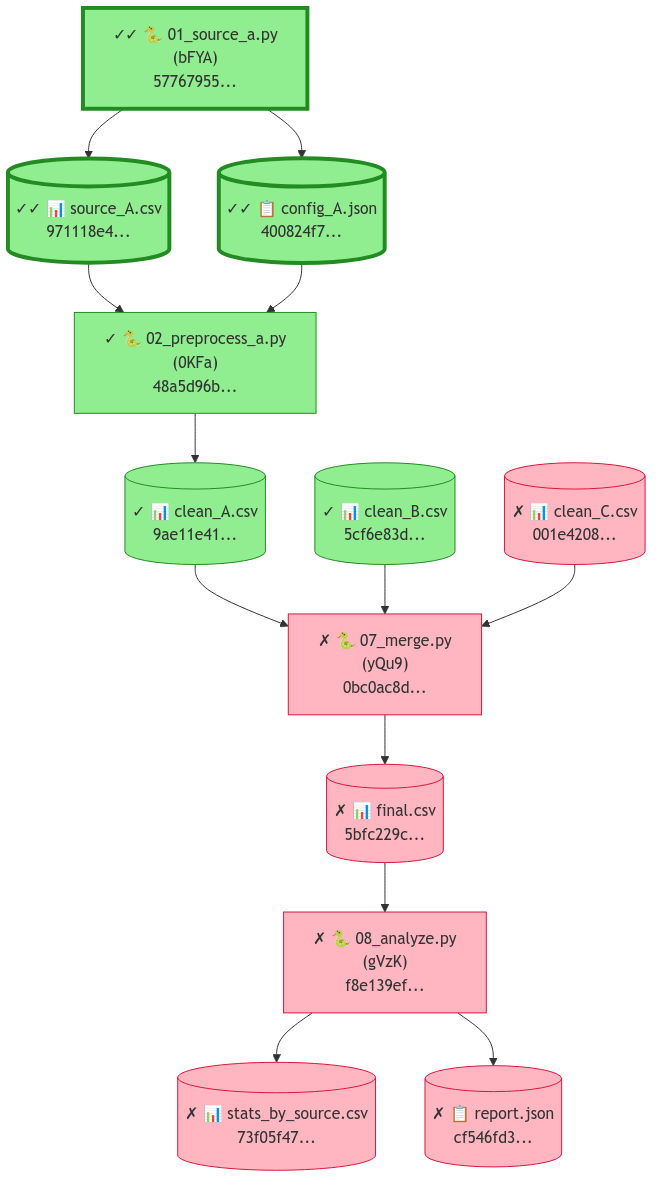
Figure 2. Clew verification DAG -- green nodes are verified (hash match), red nodes have mismatches. Each node shows its SHA-256 hash prefix.
CLI Commands
scitex --help-recursive # Show all commands
scitex scholar search "topic" # Search literature
scitex scholar fetch "10.1038/..." # Download paper by DOI
scitex stats recommend # Suggest statistical tests
scitex clew status # Project verification overview
scitex clew dag --claims # Verify all manuscript claims
scitex audio speak "Analysis complete" # Text-to-speech
scitex notification alert "Job finished" # Multi-backend notification
scitex template clone research my_proj # Scaffold a project
scitex dev versions # Check ecosystem versions
scitex mcp list-tools # List all MCP tools (293)
MCP Server (293 tools across 23 modules)
Turn AI agents into autonomous researchers via MCP.
| Category | Tools | Category | Tools | Category | Tools | ||
|---|---|---|---|---|---|---|---|
| plt | 73 | crossref | 15 | io | 5 | ||
| cloud | 50 | dev | 13 | template | 4 | ||
| writer | 38 | introspect | 12 | openalex | 4 | ||
| scholar | 22 | stats | 10 | linter | 3 | ||
| clew | 9 | dataset | 8 | social | 3 | ||
| project | 6 | notify | 5 | tunnel | 3 | ||
| docs | 4 | ui | 2 | usage | 2 |
{"mcpServers": {"scitex": {"command": "scitex", "args": ["mcp", "start"],
"env": {"SCITEX_ENV_SRC": "${SCITEX_ENV_SRC}"}}}}
Configuration
cp -r .env.d.examples .env.d # 1. Copy examples
$EDITOR .env.d/ # 2. Edit credentials
source .env.d/entry.src # 3. Source in shell
SciTeX Ecosystem
scitex-cloud is a self-hosted web application that serves as a collaborative research workspace — with a built-in Writer, Scholar, and App Store where researchers build custom tools using scitex-app SDK and scitex-ui components, then share them with the community. A live instance is hosted at scitex.ai.
Full Ecosystem (17 packages)
| Package | Module | Description |
|---|---|---|
| scitex-clew | stx.clew | SHA-256 hash-chain DAG for provenance |
| scitex-io | stx.io | Unified file I/O (30+ formats) |
| scitex-stats | stx.stats | Publication-ready statistics |
| figrecipe | stx.plt | Publication-ready matplotlib figures |
| scitex-writer | stx.writer | LaTeX manuscript compilation |
| scitex-scholar | stx.scholar | Literature management |
| scitex-notification | stx.notification | Multi-backend notifications |
| scitex-audio | stx.audio | Text-to-speech and audio |
| scitex-dev | stx.dev | Developer tools, ecosystem management |
| scitex-linter | stx.linter | AST-based code pattern checking |
| scitex-dataset | stx.dataset | Scientific datasets |
| scitex-cloud | stx.cloud | Self-hosted research platform |
| scitex-app | stx.app | Runtime SDK for research apps |
| scitex-ui | stx.ui | React/TS frontend components |
| crossref-local | stx.scholar | Local CrossRef (167M+ papers) |
| openalex-local | stx.scholar | Local OpenAlex (250M+ works) |
| socialia | stx.social | Social media (Twitter, LinkedIn) |
Four Freedoms for Research
- The freedom to run your research anywhere -- your machine, your terms.
- The freedom to study how every step works -- from raw data to final manuscript.
- The freedom to redistribute your workflows, not just your papers.
- The freedom to modify any module and share improvements with the community.
AGPL-3.0 -- because research infrastructure deserves the same freedoms as the software it runs on.